Faculty of Science Doctoral Training Centre in Artificial Intelligence
“The Faculty of Science AI DTC is a new initiative by the University of Nottingham to train future researchers and leaders to address the most pressing challenges of the 21st Century through foundational and applied AI research on a cohort basis. The training and supervision will be delivered by a team of outstanding scholars from different disciplines cutting across Arts, Engineering, Medicine and Health Sciences, Science and Social Sciences.
The Faculty of Science will invite applications from Home students for fully-funded PhD studentships to carry out multidisciplinary research in the world-transforming field of artificial intelligence. The PhD students will have the opportunity to:
- Choose from a wide choice of AI-related multidisciplinary research projects available, working with world-class academic experts in their fields;
- Benefit from a fully-funded PhD with an attractive annual tax-free stipend;
- Join a multidisciplinary cohort to benefit from peer-to-peer learning and transferable skills development.”
Studentship information
| Entry requirements |
Minimum of a 2:1 bachelor's degree in a relevant discipline to the research topic (please consult with the potential supervisors), and a strong enthusiasm for artificial intelligence research. Studentships are open to home students only |
| Start date |
1st October 2026 |
| Funding |
Annual tax-free stipend to cover living costs based on the UKRI rate (£21,805 for 2026/27), Home tuition fee, and £3000 p.a. Research Training Support Grant. |
| Duration |
3.5 years (42 Months) |
| Application deadline |
The deadline to have completed and submitted your application to NottinghamHub is Sunday 19th April 2026.
|
For information on how to apply click here
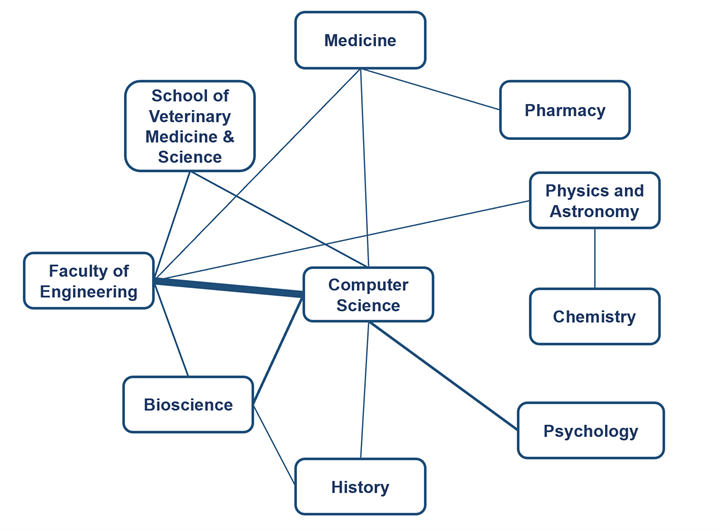
Rooted in the exceptional research environments of our Schools/Faculties at the University of Nottingham, the fifth cohort of the AI DTC will be organised around AI DTC will be organised around 13 multidisciplinary research topics. Research topics marked with (*) have strong support through match funding from industry or School sponsors, or grants.
It is important that you identify a research topic aligned with the expected skill set, your background and particular areas of interest. You will need to obtain support from the supervisors associated with your research topic choice before submitting your official application. You can do this by exploring the research projects below and contacting the main supervisor of the project that is of interest to you, directly, to discuss the further details and to arrange an interview as appropriate. In your PhD studentship application, you will be asked to provide your CV, and a personal statement including a research/project topic from the following list and explaining why you are interested in that research/project topic and your motivation for doing a PhD, and the names of the supervisors you have support from. We encourage applicants to complete the personal statement in their own words based on their background and experience. Please follow the instructions above on how to apply.
AI for additive manufacture of complex flow devices (Engineering, Computer Science)*
Medicines shortages have reached record highs in recent years due to adherence to logistically complex, labour intensive and inefficient batchwise manufacturing. High energy consumption and use of hazardous solvents cause environmental concerns, with long supply chains leading to extensive carbon footprints. This studentship will be part of a broader vision to leverage the latest advances in 3D printing and AI-driven design optimization to enable on-demand medicines manufacture that is highly cost-efficient and sustainable.
Building on our previous work the student will exploit recent advances in generative AI, including diffusion‑based models. They will optimize a) Reactor geometry to improve mixing and catalyst accessibility; b) Process control parameter data (flow rate, temperature etc.), to enable highly efficient product synthesis. The student will generate large computational model datasets, validated against the provided latest experimental data, to enable effective training and benchmarking of the AI models. Design optimization will be driven by performance metrics, guided by industrial partners from aligned grants, including Johnson Matthey and GSK. The project will equip the student with a broad array of advanced computational and AI-based skills, experience of applying those methods to optimize real-life biotechnology, industrial contacts and interdisciplinary knowledge suitable for a career in academia or industry.
The studentship is aligned with several current priority and proposed research streams within the Faculties of Computer Science and Engineering, ensuring PDRA expertise will be available to support the student. This includes the current large EPSRC Programme Grant ‘Dial Up’ (PG-‘Dial Up’ - EP/W017032/1), where one of three product objectives is a 3D printed flow reactor. The broad and thriving environment of students, researchers and academics will provide the candidate with the support, mentorship, skills & training to become an independent expert in AI optimization of 3D printed technologies with applications related to medicines synthesis and healthcare technology.
We are seeking an enthusiastic and motivated candidate with a strong quantitative background and a passion for AI driven scientific discovery, ideally with interest in geometric modelling, 3D structures, or design automation.
Essential skills include:
- Degree in Computer Science, Engineering, Artificial Intelligence, Applied Mathematics, or a related discipline.
- Strong programming skills.
- Familiarity with machine learning, especially deep learning frameworks.
- Good understanding of algorithms, numerical methods, or scientific computing.
- Desirable skills include:
- Experience with generative models
- Knowledge of (surrogate) optimisation algorithms, or (meta)heuristics.
- Background in computational fluid dynamics (CFD) or simulation based modelling (not essential; training will be provided).
- Familiarity with high performance computing (HPC) environments.
Supervisors: Dr. Simon Attwood (Faculty of Engineering), Prof. Ricky Wildman (Faculty of Engineering), Prof. Ender Ozcan (School of Computer Science), Dr. Mirco Magnini (Faculty of Engineering).
For further details and to arrange an interview please contact Dr. Simon Attwood (Faculty of Engineering).
AI-Driven Cancer Diagnosis from Imaging Mass Spectrometry Data (Pharmacy, Medicine)*
Early and accurate cancer diagnosis saves lives, yet current diagnostic approaches struggle to capture the full molecular complexity of tumours. Imaging mass spectrometry can reveal this hidden chemical information directly from biopsy samples, however, the resulting datasets are high-dimensional, complex, and difficult to interpret using conventional analytical methods.
This project is at the interface of artificial intelligence, biomedical data science, and cancer research, developing AI-based diagnostic and prognostic tools that have clear translational potential. AI and machine-learning methods will be developed to analyse imaging mass spectrometry data from cancer biopsies. A range of correlation, classification, and pattern-recognition methods will be explored, with particular emphasis on neural-network-based approaches.
In addition to computational analysis, the project will include experimental data generation. The student will receive full training in the operation of a secondary ion mass spectrometer and in associated data-processing workflows. This integrated approach will ensure a strong understanding of both the experimental origin of the data and the AI methods used for analysis.
Supervisors: Supervisors: Andrew Hook (School of Pharmacy), Grazziela Figueredo (Faculty of Medicine), Kenton Arkill (Faculty of Medicine), Cathy Merry (Faculty of Medicine), Madhumita Dandapani (Faculty of Medicine).
For further details and to arrange an interview please contact Dr. Andrew Hook (School of Pharmacy).
AI-Enhanced Conversational Support for People Living with Dementia (Engineering, Computer Science, Medicine)*
Communication difficulties, including problems with speech and language, are common among people living with dementia and often worsen as the condition progresses. These difficulties can significantly impact everyday interactions with family members, carers, and professionals, limiting social participation and engagement and reducing independence. Supporting communication is therefore central to maintaining social engagement, inclusion, and quality of life for people with dementia.
Evidence shows that people with dementia can and do use technology, particularly when it is co-designed to be accessible, meaningful, and responsive to their needs. The development of assistive technologies for people with dementia has accelerated rapidly in recent years, creating new opportunities to support communication in innovative ways.
This PhD project aims to develop an AI-enabled communication support (mobile) tool that assists people with dementia who experience difficulties with speech and language. The tool will predict likely next words during speech and, when a person becomes stuck, offers optional prompts, such as suggested words or visual cues, to support expression without taking control away from the user. Use of the tool will be aimed at maintaining good communication relationships between dyads (a person with dementia and their carer, family member or friend).
The primary focus is the development and optimisation of the underlying AI technology. However, a key aspect of this work is to co-develop the tool with people with dementia and carers, ensuring the tool is ethically grounded, usable, and aligned with real-world communication needs. This PhD would therefore suit a candidate who is interested in developing AI tools as well as working with end users and other stakeholders to achieve a practical solution.
Supervisors: Dr. Michael Craven (Engineering), Dr. Armaghan Moemeni (School of Computer Science), Dr. Esther Loseto-Gerritzen (School of Medicine).
For further details and to arrange an interview please contact Dr. Michael Craven (Engineering).
Autonomous Robotic Soil Pathogen Detection Using AI-Driven Field Exploration and Sampling (Computer Science, Biosciences)
This PhD project will develop an AI-enabled robotic system for autonomous soil pathogen detection, addressing a critical challenge in sustainable agriculture and food security. The project will integrate a quadruped robot platform (e.g. Unitree Go2, Boston Dynamics Spot) with advanced manipulation, perception, and decision-making capabilities to autonomously explore agricultural fields, collect soil samples, and interface with state-of-the-art soil pathogen detection technologies developed at the University of Nottingham’s School of Biosciences. The research will focus on enabling robust field deployment through AI-driven exploration and sampling strategies, adaptive manipulation for soil interaction, and effective human–robot interaction (HRI) to support collaboration with farmers and soil scientists. The robot will learn where, when, and how to sample soil by combining prior agronomic knowledge, real-time sensor data, and historical pathogen datasets. The long-term vision is a deployable robotic assistant that supports early pathogen detection, reduces manual labour, and enables precision intervention.
Applicants are expected to have a strong background in computer science, robotics, or a related discipline, with interests in artificial intelligence, machine learning, autonomous systems, or human–robot interaction. Experience in robotics, field robotics, or data-driven modelling is desirable, but training will be provided across disciplines. Interest in precision crop protection and/or diagnostics at the point of care (field) is highly desirable.
Key Scientific Questions
- How can autonomous robots robustly manipulate and interact with soil in unstructured agricultural environments?
- How can heterogeneous data sources (robot sensory data, spatial field data, and biological soil detection outputs) be fused to guide decision-making in situ?
- How can AI-driven exploration and reinforcement/active learning strategies optimise soil sampling for early pathogen detection under real-world field constraints?
- How can learning-based robotic systems generalise across fields, crops, and environmental conditions?
- What forms of human–robot interaction best support trust, interpretability, and effective collaboration between robotic systems, farmers, and soil scientists?
Supervisors: Dr. Ayse Kucukyilmaz (School of Computer Science), Prof. Rumiana Ray (School of Biosciences), Dr. Aly Magassouba (School of Computer Science)
For further details and to arrange an interview please contact Dr. Ayse Kucukyilmaz (School of Computer Science).
Developing AI tools to identify septic shock from photoplethysmography data (Computer Science, Engineering)
There are 245,000 cases of sepsis/year in the UK. It kills more than breast, bowel and prostate cancer combined. Globally there are 48.9 million cases resulting in 11 million deaths. Better methods are urgently needed to identify and guide treatment of sepsis. Our vision is to transform the way clinicians assess and manage patients with sepsis through a novel digital device with embedded AI-algorithms that continuously monitors the microcirculation non-invasively using photoplethysmography (PPG). PPG is the technique used in pulse oximeters and fitness monitoring devices. This PhD will investigate AI approaches to identify changes in the PPG signal that correspond to sepsis and septic shock states; and test the hypothesis that these signals can be used to detect early signs of sepsis. This project utilises sensor data acquired using instruments developed by our research team. Although most of the PhD will focus on the development of tools to analyse the data, students should have an interest in making measurements on human subjects.
Supervisors: Lucas Fonseca (School of Computer Science), Steve Morgan (Faculty of Engineering)
For further details and to arrange an interview please contact Dr. Lucas Fonseca (School of Computer Science).
Developing an AI application to quantify and classify tics in tic disorders (Psychology, Computer Science)*
Tourette syndrome is a neurological condition of childhood onset that is characterised by the occurrence of vocal and motor tics. Tics are involuntary, abrupt, recurrent, and non-rhythmic movements or vocalisations. TS affects ~1-2% of children and adolescents, is the most common form of movement disorder in children and will often have a negative impact on a child’s educational and social development, their quality-of-life, and their physical health. Accurate diagnosis and clinical management of tic disorders requires clinical expertise, in particular the ability to correctly classify and quantify tics. Currently, the most widely used method for assessing tic severity is the Yale Global Tic Severity Scale (YGTSS) which has a number of known problems and lacks objective precision. This project would develop a robust AI (machine learning) approach to the problem of tic classification and quantification based upon the analysis of video data.
Supervisors: Prof. Stephen Jackson (School of Psychology), Dr. Alexander Turner (School of Computer Science)
For further details and to arrange an interview please contact Prof. Stephen Jackson (School of Psychology).
Developing computer vision approaches to uncover new horticultural insight from lost knowledge (Computer Science, Biosciences, History, Manuscripts and Special Collections)
In this PhD you will develop cutting edge AI approaches to understanding historical manuscripts. Specifically, we wish to uncover new gardening (horticultural) insight by developing approaches to retrieve information about how gardens were developed and managed in the past. Such understanding can lead to modern day insights (new understandings of peat free soil, low-input growth, grafting, etc.) based on historical knowledge which is lost in the past. By developing modern day, computational approaches to help with searching these real-world documents, we can hope to uncover new information which will be valuable for modern horticulture.
This PhD is highly interdisciplinary. You will be developing computational approaches to understanding manuscripts held at the University of Nottingham’s Manuscripts and Special Collections (MSC) facility in Nottingham. This collection holds over 3.5 million physical documents and artefacts, including substantial collections specifically on horticulture, farming and gardening in Nottinghamshire and the UK. You will work with the manuscripts team, alongside biologists with a deeper understanding of plant science and advice from historians, to develop new ways to digitise, search and understand this data using modern approaches to AI, including computer vision and large language models. In addition, you will be working with material used by external organisations such as the National Trust who collaborate with MSC, to utilise collections relating to sites they care for, such as Clumber Park in North Nottinghamshire. Training will be available for working with the manuscripts, including how to digitise required material, both using specialist equipment at MSC and using handheld devices such as smartphones. You will be supported as part of the Computer Vision Lab at the University of Nottingham (Jubilee Campus), as well as through this DTC cohort.
A background in computer programming is essential, as you will be developing machine learning techniques and building software tools. You should also have an interest in biology and history. You do not need to be an expert in computer vision, though some experience would be a benefit. You will work with people from multiple disciplines, so good communication skills are essential.
Supervisors: Prof. Andrew French (School of Computer Science/Biosciences), Dr. Amanda Rasmussen (School of Biosciences), Dr. Michael Pound (School of Computer Science), Dr. Charlotte May (Libraries Manuscripts and Special Collections), Dr. Richard Gaunt (School of History)
For further details and to arrange an interview please contact Prof. Andrew French
Harnessing adaptation in natural materials for better engineered products (Veterinary Medicine and Science, Engineering, Computer Science)*
The natural world contains an astounding variety of organisms well-suited to their environments. From the micron-scale cellular structure of lightweight bird bones to the gradated meter-scale structure of the most resilient trees, natural materials have evolved to solve a range of challenges exceedingly well. These challenges are present in engineering too, but we are not able yet to fully exploit the solutions offered by nature. This project seizes on this opportunity with a combination of high-resolution 3D scanning, machine learning and digital manufacturing. We will employ a generative deep-learning method developed by our team in a prior project, alongside additive manufacturing methods uniquely capable of reproducing the inner structure of natural materials. The primary goal of the project will be the acquisition of a high quality data set of 3D models of natural materials using X-ray CT, focussing first on bone tissue. This will be used to train a generative AI model to create new structures with tailorable internal topology and mechanical properties. Finally, the project will demonstrate the manufacturability of the new bone architectures via additive manufacturing.
Candidates should hold a degree in computer science or the physical sciences, should have good programming skills (Python/C++) and would ideally have knowledge and/or experience of the use of AI in scientific research. The student will receive expert guidance and support from the Faculty of Engineering, School of Veterinary Medicine and Science, School of Computer Science, and the Hounsfield Facility.
Supervisors: Catrin Rutland (School of Veterinary Medicine and Science), Ian Maskery (Faculty of Engineering), Craig Sturrock (Faculty of Engineering), Victoria James (School of Veterinary Medicine and Science), Ender Ozcan (School of Computer Science)
For further details and to arrange an interview please contact Prof. Catrin Rutland
Interpretable AI for Discovering Structure–Function Rules in Submerged Fungal Fermentation (Engineering, Biosciences)
Filamentous fungal fermentation underpins a multi-billion-pound global biomanufacturing sector, spanning food, enzymes, chemicals and alternative proteins, and is expected to grow rapidly as sustainable fermentation scales. However, these systems remain difficult to optimise because fungal growth can adopt very different mycelial structures (e.g. pellets, clumps, dispersed hyphae) in response to small changes in process conditions. This PhD will develop interpretable AI methods that move beyond prediction to discover mechanistic structure–function rules from multimodal datasets, enabling improved understanding and control of morphology-driven performance in fungal fermentation.
In Year 1, the student will focus on laboratory experimentation, cultivating filamentous fungi under controlled conditions, systematically perturbing operating parameters, and collecting microscopy/morphology, sensing, and biochemical data while building strong foundations in machine learning through formal training courses, hands-on data analysis, and close shadowing of a senior PhD researcher to develop practical skills in data preprocessing, model development, and interpretation. In Year 2, the student will develop and refine interpretable AI models (e.g. symbolic regression and causal inference) to extract mechanistic structure–function relationships. In Year 3, AI predictions will guide new targeted fermentation experiments and be tested through an industrial placement, applying the developed framework to real fermentation processes.
This PhD uniquely equips the student with both experimental lab and advanced machine learning skills, producing a rare hybrid researcher fluent in biomanufacturing and AI. Applicants with either machine learning or biological, engineering, chemical manufacturing backgrounds are encouraged as any missing expertise will be developed during the PhD.
Supervisors: Oliver Fisher (Chemical and Environmental Engineering), Asma Ahmed (Chemical and Environmental Engineering), Vincenzo di Bari (School of Biosciences)
For further details and to arrange an interview please contact Dr. Oliver Fisher (Chemical and Environmental Engineering)
Machine Learning Density Functionals from Quantum Computing (Chemistry, Physics & Astronomy)*
Data is more valuable than oil, so it has been said. Quantum computing offers new unusual datasets thereby presenting new opportunities for AI approaches. Quantum computing is raising the prospect of calculations on a hardware architecture that matches the inherent nature of quantum chemistry electronic structure calculations and with it the opportunity to capture some of the inherent physics, albeit with the noise associated with near-term quantum devices. This in turn offers an exciting new dataset from which it will be possible to use machine learning to train a more accurate functional for use in density functional theory. In collaboration with Phasecraft, a leading quantum algorithms company, this project will explore the generation of new quantum computing datasets and the development of machine learning techniques to utilize the datasets to train improved density functionals for use in quantum chemical electronic structure calculations.
Applicants should have, or expected to achieve, at least a 2:1 Honours degree (or equivalent if from other countries) in Chemistry, Physics, Mathematics, Computer Science or Natural Sciences or a related subject. A MChem/MSc-4-year integrated Masters, a BSc + MSc or a BSc with substantial research experience will be highly advantageous. Experience in computer programming will be essential.
Supervisors: Jonathan Hirst (School of Chemistry), Katherine Inzani (School of Chemistry), Adam Gammon-Smith (School of Physics & Astronomy)
For further details and to arrange an interview please contact Prof. Jonathan Hirst (School of Chemistry)
Physics-Informed Multi-Agent AI for Cooperative Formation-Flying Orbital Transfers (Engineering, Computer Science)
Satellites operating in coordinated groups require autonomous, fuel-efficient, and safe reconfiguration strategies to realise next-generation Earth observation, navigation, and in-orbit services missions. This project will address these challenges by developing novel AI-driven methods for cooperative orbital transfers across satellite formations. The research will combine multi-agent reinforcement learning (MARL) with physics-informed modelling of relative orbital dynamics to enable distributed decision-making among spacecraft under realistic perturbations (e.g. J₂, drag, solar radiation pressure) and communication constraints. The PhD student will design learning frameworks that explicitly incorporate collision avoidance, fuel efficiency, and real-time adaptability, moving beyond centralised optimisation to scalable, agent-level autonomy.
The project will be supported by an interdisciplinary supervisory team spanning Engineering and Computer Science, providing multidisciplinary training in AI, control theory, optimisation, and astrodynamics. The student will benchmark AI methods against classical transfer and formation control techniques and validate solutions using high-fidelity simulations and hardware-in-the-loop testbeds. This work is ideal for applicants with interests in AI, autonomous systems, and space sciences.
Applicants should hold (or expect to obtain) a first-class undergraduate degree (or equivalent), or a distinction at postgraduate level, in Aerospace Engineering, Mechanical Engineering, Electrical Engineering, Computer Science, Physics, or a closely related discipline. Essential skills include strong mathematical foundations (e.g. linear algebra, optimisation, and dynamical systems), proficiency in Python, and experience or a strong interest in both machine learning and dynamical/control systems. Desirable experience includes reinforcement learning, multi-agent systems, orbital mechanics, and numerical simulation. The candidate should be motivated to work at the interface of AI and space systems and be comfortable working across Engineering and Computer Science disciplines.
Supervisors: Nishanth Pushparaj (Faculty of Engineering), Chantal Cappelletti (Faculty of Engineering), Nikhil Deshpande (School of Computer Science)
For further details and to arrange an interview please contact Dr. Nishanth Pushparaj (Faculty of Engineering)
The Metabolic Cost of Training: Bridging Synaptic Plasticity Theory and Data-Centric Efficiency in Deep Learning (Psychology, Computer Science)
The rapid growth of deep learning has come at an extraordinary environmental and computational cost, yet the standard training paradigm remains remarkably unchanged. Every sample is passed through the network, and every synapse is updated, on every epoch. Recent work has begun to challenge both halves of this independently. Progressive Data Dropout has shown that progressively reducing the training set across epochs can cut effective training epochs by up to 90% while actually improving accuracy. Separately, research grounded in neuroscience has demonstrated that restricting synaptic updates to only the most informative weights, motivated by the metabolic cost of learning in biological systems, can reduce the energetic burden of training by orders of magnitude. What has not yet been explored is what happens when these two ideas are unified under a single energy-aware framework. This PhD will develop principled methods for jointly optimising which data and which weights are updated during training, using metabolic energy as the governing design constraint. The student will build on an established energy model for synaptic plasticity to derive theoretically grounded training schedules, and will evaluate these across a range of architectures (CNNs, Vision Transformers, and language models) on standard benchmarks through to large-scale settings.
The project sits at the intersection of machine learning, computational neuroscience, and cognitive science, and the student will work closely with both supervisors to move between these perspectives. We are looking for a candidate with a strong foundation in either machine learning or mathematical/computational neuroscience, demonstrable programming experience (Python/PyTorch), and the curiosity to work across disciplinary boundaries. A background in optimisation theory or an interest in the energy and sustainability implications of AI would be particularly welcome.
Supervisors: Prof. Mark van Rossum (School of Psychology), Dr. Shreyank Narayana Gowda (School of Computer Science)
For further details and to arrange an interview please contact Prof. Mark van Rossum (School of Psychology)
Unsupervised Discovery in Atomic-scale Spectroscopy (Physics & Astronomy, Computer Science)
Scanning probe microscopy uniquely combines ultrahigh-resolution imaging with targeted, site-specific manipulation of matter right down to the single-atom, single-molecule, and single-chemical-bond levels. This makes it arguably the most powerful tool available for configuring and controlling condensed matter. Beyond imaging and manipulation, scanning probe microscopes provide powerful spectroscopic access to matter at the same fundamental length scales and with exceptional energy resolution. By measuring electronic, vibrational, and chemo-mechanical responses as a function of parameters controlled by the microscopist (including voltage, tip-sample distance, electrostatic or magnetic field strength, etc..), rich multidimensional maps of atomic, molecular, and/or nanoscale properties can be acquired. At the same time, the very sensitivity that makes probe spectroscopy so powerful also makes its interpretation challenging: measured spectra very often reflect a complex convolution of sample properties, tip structure, and experimental conditions.
This PhD project will use unsupervised and self-supervised machine learning to explicitly disentangle these contributions by analysing large ensembles of spectra acquired under varying conditions. By learning latent representations that separate tip-dependent and sample-dependent features, the project aims to identify changes in tip state, recover intrinsic sample fingerprints, and reduce systematic ambiguity in spectroscopic interpretation. This will involve the development and application of unsupervised representation-learning techniques, such as autoencoders and related generative models, together with clustering and dimensionality-reduction methods tailored to spectroscopic data. Ultimately, we aim to fundamentally change how information is extracted from atomic-scale spectroscopy so that the data speaks for itself.
For more on the scanning probe microscopy-machine learning interface, see:
- OM Gordon and PJ Moriarty, Mach. Learn.: Sci. Technol. 1 023001 (2020)
- U Pratiush, et al., Digital Discovery, 4, 252 (2025)
For videos of the ultrahigh vacuum, low temperature scanning probe microscopes (here at UoN) that will generate the spectra to be analysed in this project, see:
Supervisors: Prof. Philip Moriarty (School of Physics & Astronomy), Dr. Michael Pound (School of Computer Science), Dr. Brian Kiraly (School of Physics & Astromomy)
For further details and to arrange an interview please contact Prof. Philip Moriarty (School of Physics & Astronomy)
Research topics marked with (*) have strong support through match funding from industry or School sponsors, or grants
Further information
For further enquiries, please contact Professor Ender Özcan - School of Computer Science